-
News
Automatic quantification of cardiomyocyte dimensions and connexin 43 lateralization in fluorescence images
Oct 20, 2020
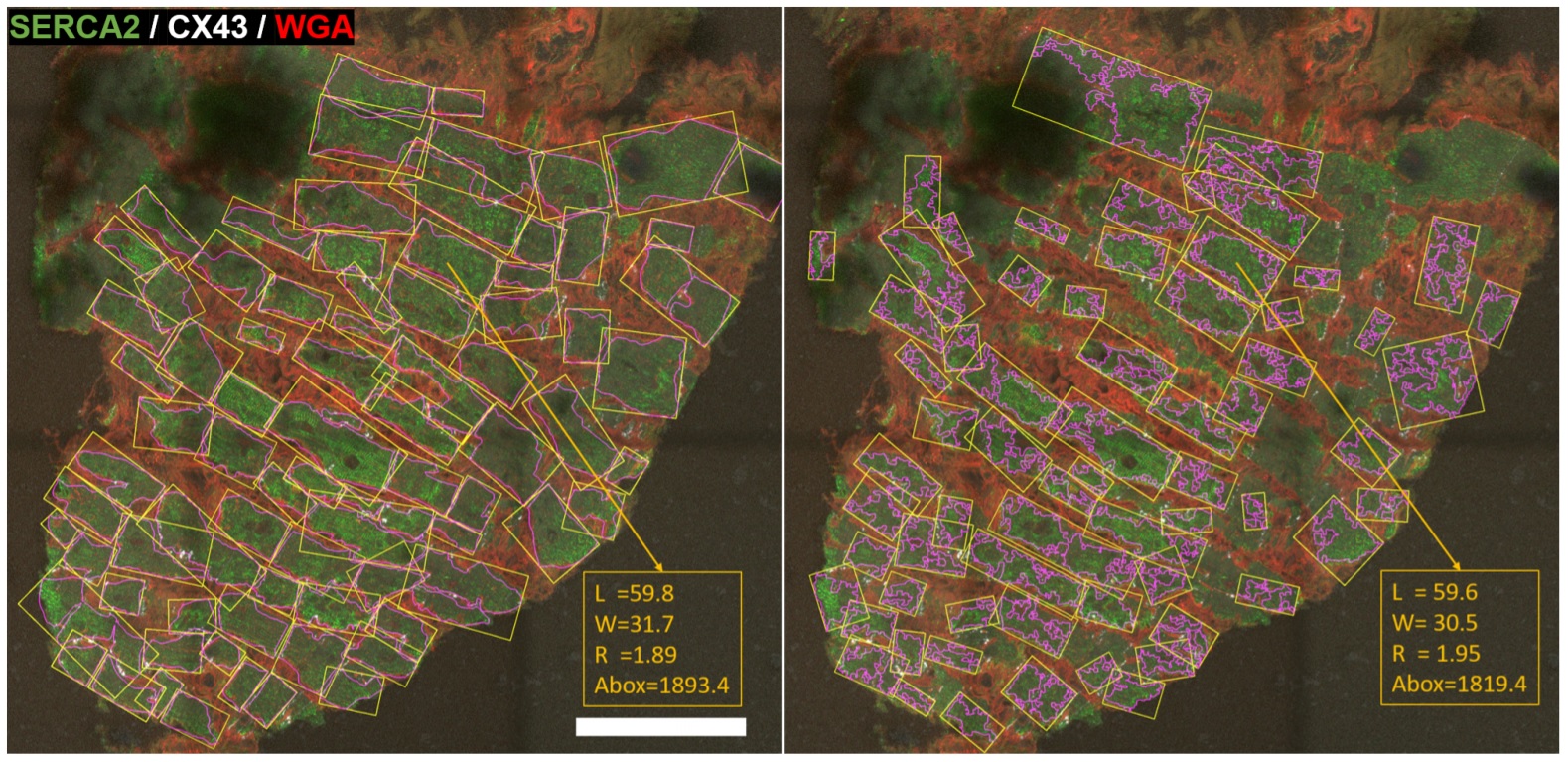
Cardiomyocytes’ geometry and connexin 43 (CX43) amount and distribution are structural features that play a pivotal role in electrical conduction. Their quantitative assessment is of high interest in the study of arrhythmias, but it is usually hampered by the lack of automatic tools. In this work, we propose a software algorithm (Myocyte Automatic Retrieval and Tissue Analyzer, MARTA) to automatically detect myocytes from fluorescent microscopy images of cardiac tissue, measure their morphological features and evaluate the expression of CX43 and its degree of lateralization.
The proposed software is based on the generation of cell masks, contouring of individual cells, enclosing of cells in minimum area rectangles and splitting of these rectangles into end-to-end and middle compartments to estimate CX43 lateral-to-total ratio. Application to human ventricular tissue images shows that mean differences between automatic and manual methods in terms of
cardiomyocyte length and width are below 4 µm.
The percentage of lateral CX43 also agrees between automatic and manual evaluation, with the interquartile range approximately covering from 3% to 30% in both cases. MARTA is not limited by fiber orientation and has an optimized speed by using contour filtering, which makes it run hundreds of times faster than a trained expert. Developed for CX43 studies in the left ventricle, MARTA is a flexible tool applicable to morphometric and lateralization studies of other markers in any heart chamber or even skeletal muscle. This open-access software is available online.
A. Oliver-Gelabert, L. García-Mendívil, J.M. Vallejo-Gil, P. C. Fresneda-Roldán, K. Andelová, J. Fañanás-Mastral, M. Vázquez-Sancho, M. Matamala-Adell, F.
Sorribas-Berjón, C. Ballester-Cuenca, N. Tribulova, L. Ordovás, E.R. Diez, E. Pueyo (2020). Automatic quantification of cardiomyocyte dimensions and connexin
43 lateralization in fluorescence images. Biomolecules. 2020, 10(9), 1334
Full Article: https://doi.org/10.3390/biom10091334